A more comprehensive epigenomic map of the dog captures significant similarities with the human epigenome paving the way for comparative biology and medicine
Researchers have successfully mapped the dog epigenome and created a high-quality reference map driving new understanding and opening new avenues for functional genomics research in dogs and comparative studies with humans and other species. The pioneering work, appearing in the July 5 edition of Science Advances, provides an in-depth annotation of epigenomic marks which are modifications on DNA that can influence gene activity without changing the underlying genetic code.
The remarkable diversity of dog breeds, developed within a relatively short span of tens of thousands of years, holds valuable insights into complex morphological and behavioral traits, genetic disorders, and even diseases like cancer. By leveraging state-of-the-art technologies, researchers can delve deep into the genetic makeup of dogs, unraveling their inner workings and, in turn, enhancing our understanding of human health and diseases. This has been true since the publication of the first draft of the dog genome in 2005, a monumental achievement that provided a blueprint of encoded genetic information in the DNA of dogs. However, unlike in the case of humans and mice, dog genomics has long been lacking a reference epigenome, leaving a significant gap in our understanding of their gene regulation and the impact of epigenetic modifications.
Dogs, often regarded as both cherished companions and essential livestock animals, have coexisted with humans for thousands of years, sharing a symbiotic relationship marked by a shared environment, diet, lifestyle, and exposure to infectious agents. Despite this close association, there has been a noticeable absence of research focusing on how both humans and dogs are influenced by their shared surroundings, leaving a critical knowledge gap in understanding the environmental factors that impact their health and well-being.
Understanding the effects of environmental factors necessitates crucial understanding of the dog epigenome. Unlike the genome, which remains relatively insensitive to environmental influences, the epigenome provides valuable insights into the impact of environmental factors. For instance, studies on twins, who possess identical genomes but lead distinct lifestyles, have uncovered remarkable differences in their epigenetic profiles, highlighting the role of the epigenome in reflecting environmental influences. By delving into the dog epigenome, scientists can unravel the intricate relationship between environmental factors and gene regulation, shedding light on how dogs respond and adapt to their surroundings.
Dogs are also known for their accelerated biological clocks and shorter lifespans compared to humans. For these reasons, dogs possess the ability to serve as sentinels, rapidly reacting to environmental risk factors and alerting humans to potential dangers. In this context, the dog epigenome emerges as a highly sensitive factor, offering valuable insights. A comprehensive understanding of both the dog genome and epigenome holds paramount importance for advancing biomedical sciences, benefiting animals and humans alike.
To tackle these challenges, a team led by Professor Je-Yoel Cho from the College of Veterinary Medicine and Comparative Medicine Disease Research Center (CDRC) at Seoul National University has unveiled for the first time a reference annotation of the dog epigenome. They meticulously examined 11 major dog tissues, including the cerebrum, cerebellum, mammary gland, lung, liver, stomach, spleen, pancreas, kidney, colon, and ovary. Through the generation and analysis of diverse epigenomic data, they constructed of the world's first comprehensive functional annotation of the dog genome. This groundbreaking achievement now enables the interpretation of the regulatory code governing genome activity, offering fresh insights into various biological functions, gene specificity in different cells and tissues, dysregulation of gene activity due to environmental factors, and the occurrence of diseases.
Furthermore, the research team uncovered conserved and dynamic functional characteristics shared between different tissues and species. Notably, the dog epigenome exhibited a closer resemblance to the human epigenome than that of mouse, suggesting important similarities in gene regulation and potential implications for human health.
The completion of this comprehensive epigenomic map for the dog genome marks a significant milestone in our quest to understand the intricate workings of gene regulation. The findings from this pioneering research have the potential to revolutionize biomedical research, aiding in the development of targeted treatments, enhanced disease management, and improved well-being for both dogs and humans.
Professor Cho explained, "This groundbreaking epigenome map can be extensively utilized to study different dog breeds, delve into cancer and disease mechanisms, conduct comparative research across species, and significantly contribute to advancements in human life sciences,".
He added, “This work also represents a milestone for basic research in the field of veterinary medicine. This breakthrough enables researchers to unravel the impact of epigenetic modifications on gene expression and opens up new avenues for investigating the underlying mechanisms of complex diseases, advancing veterinary diagnostics, therapeutics, and personalized medicine approaches for dogs.”
This study was supported by the Bio & Medical Technology Development Program (grant no. NRF-2016M3A9B6026771), awarded as part of the Comparative Medicine initiative, and by the Science Research Center (SRC) Program (grant no. NRF-2021R1A5A1033157) under the Directorate for Basic Research in Science & Engineering, awarded as part of the Comparative Medicine Disease Research Center (CDRC) initiative, through the National Research Foundation (NRF) funded by the Korean government’s Ministry of Science and ICT.
[Image]

(L to R) Co-first author Mark Borris D. Aldonza, Ph.D. student. corresponding author Je-Yoel Cho, DVM, Ph.D. co-first author Keun Hong Son, Ph.D. student. co-first author A-Reum Nam, Ph.D. student. Seoul National University, Comparative medicine Disease Research Center (CDRC)
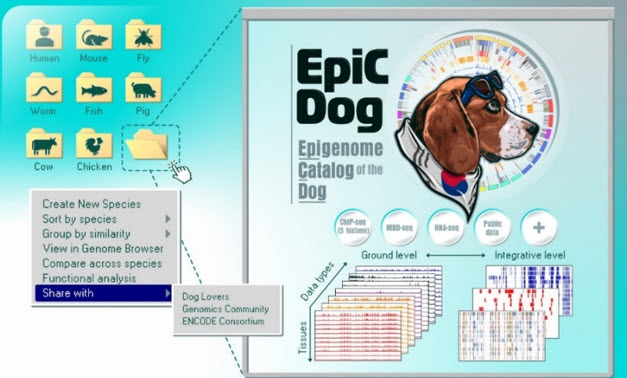
The EpiC Dog initiative.
[Journal]
Science Advances
[Article title]
Integrative mapping of the dog epigenome: reference annotation for comparative intertissue and cross-species studies
[Article publication data]
5-Jul-2023

